Calculating the Schottky barrier: Difference between revisions
No edit summary |
No edit summary |
||
| (27 intermediate revisions by 2 users not shown) | |||
| Line 1: | Line 1: | ||
The Schottky barrier is the energy barrier that forms at a metal–semiconductor junction | The [https://en.wikipedia.org/wiki/Schottky_barrier| Schottky barrier] is the energy barrier that forms at a metal–semiconductor junction. The barrier height is an important characteristic for charge transport across the junction and a critical parameter in the design of semiconductor contacts, transistors, and rectifying diodes. The p-type Schottky barrier height <math>\varphi_{\mathrm{p}}</math> describes the barrier seen by holes in the semiconductor valence band, and the n-type Schottky barrier height <math>\varphi_{\mathrm{n}}</math> describes the barrier seen by electrons in the conduction band. In {{VASP}}, the potential alignment method can be used to estimate the Schottky barrier{{cite|baldereschi:prl:1988}}{{cite|peressi:1998}}. | ||
== Required quantities == | |||
Three quantities from bulk calculations, the Fermi energy <math>E_{\mathrm{F}}</math> of the metal, and the valence-band maximum <math>E_{\mathrm{VBM}}</math> and band gap <math>E_{\mathrm{g}}</math> of the semiconductor, are needed for the calculation. In addition, the macroscopic electrostatic potential difference <math>\Delta\bar{V}</math> across the interface is extracted from a separate interface slab calculation<ref>In {{VASP}}, the <math>\mathbf{G}=0</math> Fourier component of the Hartree potential is set to zero by convention, so all single-particle eigenvalues are already referenced to the average electrostatic potential of their respective unit cell. As a consequence, <math>E_{\mathrm{F}}</math> and <math>E_{\mathrm{VBM}}</math> can be read directly from the bulk {{FILE|OUTCAR}} files without any further potential referencing, and only the interface calculation requires the electrostatic potential to be written out — to {{FILE|vaspout.h5}} via {{TAG|WRT_POTENTIAL}} (recommended) or to {{FILE|LOCPOT}} via {{TAG|LVHAR}} (legacy).</ref>. | |||
== Step-by-step instructions == | == Step-by-step instructions == | ||
{{NB|important|For the calculation of the p- and n-type Schottky barriers, energies, and potentials from different VASP calculations are compared. It is crucial to set a single value for {{TAG|ENCUT}} for all calculations.}} | |||
'''Step 1:''' Compute the Fermi energy of the metal. | |||
: After ensuring that the [[Structure optimization|bulk geometry is optimized]], a separate static calculation should be performed with a dense, Γ-centered [[KPOINTS|'''k'''-point mesh]] and {{TAG|ISMEAR|-5}} (tetrahedron method with Blöchl corrections), to get an accurate Fermi energy. It can be taken from the {{FILE|OUTCAR}} and is also easily accessible via {{py4vasp}}: | |||
: <syntaxhighlight lang="python"> | |||
{{TAG|ISMEAR|-5}} (tetrahedron method with Blöchl corrections) | |||
<syntaxhighlight lang="python"> | |||
import py4vasp as pv | import py4vasp as pv | ||
calc = pv.Calculation.from_path(" | calc = pv.Calculation.from_path("./Step1/") | ||
E_F = calc.dos.read()["fermi_energy"] | E_F = calc.dos.read()["fermi_energy"] | ||
print(f"E_F = {E_F:.4f} eV") | print(f"E_F = {E_F:.4f} eV") | ||
</syntaxhighlight> | </syntaxhighlight> | ||
'''Step 2:''' Compute the band gap and valence-band maximum of the semiconductor. | |||
: Band gaps are notoriously underestimated by standard DFT, so for accurate values, it is usually necessary to use [[:Category:DFT+U|DFT+U]], [[:Category:Hybrid functionals|hybrid functionals]], or [[:Category:GW|GW]] calculations. The level of theory to choose depends on the semiconductor of interest, and HSE06 is oftern a good default for most covalent and ionic semiconductors. | |||
: | : Generally, it is advisable to select {{TAG|BANDGAP|KPOINT}} to get more verbose output, and ideally to make a full [[:Category:Band_structure|band-structure calculation]], since the extrema of the bands are often located at high-symmetry lines. The fundamental bandgap <math>E_{\mathrm{g}}</math> and the valence-band maximum (VBM) <math>E_{\mathrm{VBM}}</math>, can be read from the {{FILE|OUTCAR}} file, or with {{py4vasp}}: | ||
:<syntaxhighlight lang="python"> | |||
import py4vasp as pv | |||
calc = pv.Calculation.from_path("./Step2/") | |||
E_g = calc.bandgap.fundamental() | |||
E_VBM = calc.bandgap.valence_band_maximum() | |||
: | print(f"E_g = {E_g:.4f} eV") | ||
print(f"E_VBM = {E_VBM:.4f} eV") | |||
</syntaxhighlight> | |||
'''Step 3:''' Build and relax an interface structure. | |||
:Building a lattice-matched metal–semiconductor interface slab is outside the scope of this page. Tools such as the <tt>CoherentInterfaceBuilder</tt> in pymatgen{{cite|pymatgen}} and the interface construction utilities in ASE{{cite|ase}} can be used to construct the initial geometry. An interface structure for a Schottky barrier calculation should satisfy the following requirements: | |||
:* ''Low lattice mismatch.'' The supercell should be chosen such that the in-plane periodicities of the two surfaces match to within ~2%, minimising artificial biaxial strain. | |||
:* ''Sufficient slab thickness.'' Each material should contain enough layers that a bulk-like region exists in the centre. | |||
:* ''No vacuum region.'' A cell with a vacuum will result in lateral strain, compressing the cell to reduce the surface energy. It is better to have two equivalent interfaces in the cell without any vacuum at all. | |||
: For the relaxation, it is usually prudent to fix the layers in the bulk like regions using [[POSCAR#poscar6|selective dynamics]] in the {{FILE|POSCAR}} file. Only a couple of layers at the interfaces should be relaxed. Depending on the material pairing and the strain in the interface, it might be beneficial to allow the cell to relax only in the direction normal to the interface plane using the {{TAG|LATTICE_CONSTRAINTS}} tag or the {{FILE|ICONST}} file to apply the contraint. | |||
'''Step 4:''' Compute the electrostatic potential of the interface and calculate the Schottky barrier heights. | |||
:Perform a static calculation with tetrahedron smearing ({{TAG|ISMEAR|-5}}) and {{TAG|PREC|Accurate}}. We need to print out the electrostatic part of the potential (disregarding the XC part) by setting {{TAG|WRT_POTENTIAL|hartree ionic}}. It is also possible to use {{TAG|LVHAR|.TRUE.}} to get the same data, but be aware that {{TAG|LVTOT}} would include the (semi-)local exchange-correlation potential <math>V_{\text{xc}}(\mathbf{r})</math>, which we want to exclude here. | |||
:The planar average <math>\bar{V}(z)</math> of the electrostatic potential is obtained by averaging the 3D potential over the in-plane (xy) grid at each <math>z</math> point (assuming the interface is located in the xy-plane). Then, a macroscopic average is computed by applying a uniform box-car convolution of window length <math>L</math>, where <math>L</math> equals one bulk primitive-cell repeat period along the interface normal. Averaging over exactly one such period removes the short-range oscillations within each atomic layer while preserving the long-range potential step across the interface. The Python script below attempts to find the optimal <math>L</math> automatically by sweeping a range of values and selecting the one that minimizes the RMS gradient of the macroscopic average in the bulk-like plateau regions of both materials simultaneously. It reads the potential from {{FILE|vaspout.h5}}, computes both averages, plots the data and reports the macroscopic electrostatic potential difference <math>\Delta\bar{V}</math>: | |||
</ | |||
:<div class="toccolours mw-customtoggle-potentialscript">'''Click to show python script'''</div> | |||
<div class="mw-collapsible mw-collapsed" id="mw-customcollapsible-potentialscript"> | |||
:<syntaxhighlight lang="python"> | |||
#!/usr/bin/env python3 | |||
""" | |||
Potential alignment script for Schottky barrier height calculations. | |||
The electrostatic potential (hartree + ionic) is read from vaspout.h5, averaged | |||
in the xy-plane to give the planar average V_bar(z), and then smoothed with a | |||
box-car macroscopic average of width L to remove short-range oscillations. | |||
dV_bar = <V_bar>_M - <V_bar>_SC is the potential offset between the metal (M) | |||
and semiconductor (SC) bulk-like regions of the slab. | |||
L is chosen automatically by minimising the worst-case RMS gradient in the | |||
two plateau regions (M and SC separately). This ensures both regions are flat, | |||
even when the two materials have different lattice periodicities along z. | |||
The plateau centres are given as fractional cell coordinates via --mc (metal) | |||
and --scc (semiconductor). A fixed plateau half-width of 0.08 * c is used. | |||
""" | |||
import argparse | |||
import matplotlib | |||
matplotlib.use("Agg") # non-interactive backend; remove if you want a GUI window | |||
import matplotlib.pyplot as plt | |||
import h5py | |||
import numpy as np | |||
from scipy.ndimage import uniform_filter1d | |||
parser = argparse.ArgumentParser(description="Compute macroscopic potential alignment.") | |||
parser.add_argument( | |||
"--mc", "--metal-center", | |||
type=float, | |||
default=0.125, | |||
metavar="F", | |||
dest="metal_center", | |||
help="Fractional z-coordinate of the metal plateau centre (default: 0.125)", | |||
) | |||
parser.add_argument( | |||
"--scc", "--sc-center", | |||
type=float, | |||
default=0.5, | |||
metavar="F", | |||
dest="sc_center", | |||
help="Fractional z-coordinate of the semiconductor plateau centre (default: 0.5)", | |||
) | |||
parser.add_argument( | |||
"--vaspout", | |||
default="./Step4/vaspout.h5", | |||
metavar="FILE", | |||
help="Path to vaspout.h5 (default: ./vaspout.h5)", | |||
) | |||
args = parser.parse_args() | |||
# -- load potential and cell geometry ------------------------------------------ | |||
with h5py.File(args.vaspout, "r") as f: | |||
hartree = f["/results/potential/hartree"][0] # (NGZ, NGY, NGX) | |||
ionic = f["/results/potential/ionic"][0] # (NGZ, NGY, NGX) | |||
lat = f["/results/positions/lattice_vectors"][:] # (3, 3) Ang | |||
c = np.linalg.norm(lat[2]) # cell length along interface normal (Ang) | |||
NGZ = hartree.shape[0] # number of grid points along z | |||
dz = c / NGZ # grid spacing (Ang) | |||
: | # -- planar average: mean over the xy-plane at each z -------------------------- | ||
V_planar = (hartree + ionic).mean(axis=(1, 2)) # shape (NGZ,) [eV] | |||
z = np.arange(NGZ) * dz # z-coordinates in Ang | |||
# -- plateau masks (used both for auto-selection and for dV_bar extraction) ---- | |||
# z_frac maps each grid point to a fractional cell coordinate in [0, 1). | |||
# For each material the mask selects a symmetric window of width 2*HALF_WIDTH | |||
# centred on the user-supplied fractional coordinate (--mc / --scc). The | |||
# distance is computed with periodic boundary conditions (minimum image), so | |||
# --mc=0.0 correctly selects both the bottom edge (z_frac < 0.08) and the | |||
# top edge (z_frac > 0.92) of the cell, which belong to the same metal region. | |||
# HALF_WIDTH = 0.08 means the window covers 16 % of the cell on each side, | |||
# which is large enough to average over several bulk-like repeat units while | |||
# staying well clear of the interface region for any reasonably converged | |||
# slab (total cell >= ~40 Ang, >= 6 layers per material). | |||
HALF_WIDTH = 0.08 | |||
z_frac = z / c | |||
: | def _periodic_mask(center): | ||
d = np.abs(z_frac - center) | |||
d = np.minimum(d, 1.0 - d) # minimum-image distance in fractional coords | |||
return d < HALF_WIDTH | |||
M_mask = _periodic_mask(args.metal_center) | |||
SC_mask = _periodic_mask(args.sc_center) | |||
# -- auto-select window length ------------------------------------------------- | |||
# Sweep L from 2*dz to c/3. For each L the score is the worst-case RMS gradient | |||
# across the two plateau regions (M and SC scored independently). Minimising the | |||
# worst case forces both regions to be flat simultaneously. | |||
L_grid = np.linspace(2 * dz, c / 3, 300) | |||
def _plateau_roughness(L): | |||
Vm = uniform_filter1d(V_planar, size=max(1, int(round(L / dz))), mode="wrap") | |||
g = np.gradient(Vm) | |||
return max(np.sqrt(np.mean(g[M_mask]**2)), | |||
np.sqrt(np.mean(g[SC_mask]**2))) | |||
scores = [_plateau_roughness(L) for L in L_grid] | |||
WINDOW_LENGTH = float(L_grid[np.argmin(scores)]) | |||
print(f"Auto-selected window length: {WINDOW_LENGTH:.2f} Ang") | |||
# -- macroscopic average: box-car convolution with period L -------------------- | |||
# uniform_filter1d computes the running mean over `size` consecutive points. | |||
# mode='wrap' enforces periodic boundary conditions, which is correct for | |||
# a slab calculation where the potential is periodic along z. | |||
n_window = int(round(WINDOW_LENGTH / dz)) | |||
V_macro = uniform_filter1d(V_planar, size=n_window, mode="wrap") | |||
# -- plot ---------------------------------------------------------------------- | |||
fig, ax = plt.subplots(figsize=(10, 4)) | |||
ax.plot(z, V_planar, lw=0.7, color="steelblue", alpha=0.5, label="Planar average") | |||
ax.plot(z, V_macro, lw=2.0, color="tomato", | |||
label=f"Macroscopic average (L = {WINDOW_LENGTH:.2f} Ang, {n_window} pts)") | |||
ax.set_xlabel("z (Ang)") | |||
ax.set_ylabel("Electrostatic potential (eV)") | |||
ax.set_xlim(0, c) | |||
ax.legend() | |||
ax.set_title("M/SC interface - electrostatic potential along interface normal") | |||
plt.tight_layout() | |||
plt.savefig("potential_alignment.png", dpi=150) | |||
print("Figure saved: potential_alignment.png") | |||
# -- extract dV_bar from bulk-like plateaus ------------------------------------ | |||
# M_mask and SC_mask are defined above; adjust --mc / --scc if the plateaus shift. | |||
V_M = V_macro[M_mask].mean() | |||
V_SC = V_macro[SC_mask].mean() | |||
delta_V = V_M - V_SC | |||
print(f"\n<V_bar>_M = {V_M:+.4f} eV") | |||
print(f"<V_bar>_SC = {V_SC:+.4f} eV") | |||
print(f"dV_bar = <V_bar>_M - <V_bar>_SC = {delta_V:+.4f} eV") | |||
</syntaxhighlight> | |||
</div> | |||
'''Step | '''Step 5:''' Compute the Schottky barrier heights. | ||
With <math>E_{\mathrm{F}}</math>, <math>E_{\mathrm{VBM}}</math>, <math>E_{\mathrm{g}}</math>, and <math>\Delta\bar{V}</math> in hand, the potential alignment difference <math>\Delta\bar{V} = \bar{V}_{\mathrm{m}} - \bar{V}_{\mathrm{sc}}</math> | :With <math>E_{\mathrm{F}}</math>, <math>E_{\mathrm{VBM}}</math>, <math>E_{\mathrm{g}}</math>, and <math>\Delta\bar{V}</math> in hand, the potential alignment difference <math>\Delta\bar{V} = \bar{V}_{\mathrm{m}} - \bar{V}_{\mathrm{sc}}</math> connects the bulk reference frames. The p-type and n-type Schottky barrier heights are then | ||
::<math> | ::<math> | ||
| Line 138: | Line 207: | ||
</math> | </math> | ||
== Example == | |||
<!--[[File:Si111_Al111_interface.png|thumbnail|120px|right|Al(111) Si(111) interface]]--> | |||
Here we will consider the Al(111)/Si(111) interface. The bulk Al calculation to determine the Fermi energy is straightforward and yields <math>E_{\mathrm{F}}=8.085</math> eV for the PBE functional with {{TAG|ENCUT|300}}, {{TAG|KSPACING|0.2}}, and {{TAG|ISMEAR|-5}}. | |||
For the Si bandgap and valence band maximum, a little more care is needed. After relaxing with PBEsol to get the correct lattice parameter, a band structure calculation with the HSE06 functional yields a bandgap of <math>E_{\mathrm{g}}=1.160</math> eV and a valence band maximum of <math>E_{\mathrm{VBM}}=5.449</math> eV. | |||
For Al(111) and Si(111), a 4×4 Al supercell matched to a 3×3 Si supercell achieves a mismatch of 0.9%. For this example, we chose 13 Al layers, and 6 Si bilayers. The starting interface distance before relaxation was set to 2.3 Å We have fully relaxed the cell, but fixed all but the 4 layers closest to the interfaces (2 Al layers, and 1 Si bi-layer for each interface) with the [[POSCAR#poscar6|selective dynamics]] {{FILE|POSCAR}} option. The relaxed structure with its 316 atoms is given here: | |||
<div class="toccolours mw-customtoggle-poscar">'''Click to show POSCAR'''</div> | |||
<div class="mw-collapsible mw-collapsed" id="mw-customcollapsible-poscar"> | |||
<pre> | |||
Al208 Si108 | |||
1.00000000000000 | |||
11.4573925481650338 0.0000000000000000 0.0000000000000007 | |||
5.7286962740825134 9.9223930078414337 0.0000000000000007 | |||
-0.0000000000000000 -0.0000000000000000 49.5333046267364523 | |||
Al Si | |||
208 108 | |||
Selective dynamics | |||
Direct | |||
0.9583333333333357 0.9583333333333357 0.2851012628001968 F F F | |||
0.9583333333333357 0.2083333333333357 0.2851012628001968 F F F | |||
0.9583333333333357 0.4583333333333357 0.2851012628001968 F F F | |||
0.9583333333333357 0.7083333333333357 0.2851012628001968 F F F | |||
0.2083333333333357 0.9583333333333357 0.2851012628001968 F F F | |||
0.2083333333333357 0.2083333333333357 0.2851012628001968 F F F | |||
0.2083333333333357 0.4583333333333357 0.2851012628001968 F F F | |||
0.2083333333333357 0.7083333333333357 0.2851012628001968 F F F | |||
0.4583333333333357 0.9583333333333357 0.2851012628001968 F F F | |||
0.4583333333333357 0.2083333333333357 0.2851012628001968 F F F | |||
0.4583333333333357 0.4583333333333357 0.2851012628001968 F F F | |||
0.4583333333333357 0.7083333333333357 0.2851012628001968 F F F | |||
0.7083333333333357 0.9583333333333357 0.2851012628001968 F F F | |||
0.7083333333333357 0.2083333333333357 0.2851012628001968 F F F | |||
0.7083333333333357 0.4583333333333357 0.2851012628001968 F F F | |||
0.7083333333333357 0.7083333333333357 0.2851012628001968 F F F | |||
0.8749353467083437 0.8749353467083437 0.2373114907195359 T T T | |||
0.8763358121166870 0.1243320939416563 0.2376670183932849 T T T | |||
0.8749353467083437 0.3751293065833128 0.2373114907195359 T T T | |||
0.8740396730492660 0.6254801634753705 0.2371083546269586 T T T | |||
0.1243320939416563 0.8763358121166870 0.2376670183932849 T T T | |||
0.1243320939416563 0.1243320939416563 0.2376670183932849 T T T | |||
0.1228899057605569 0.3749413149216736 0.2378372905376746 T T T | |||
0.1228899057605569 0.6271687793177693 0.2378372905376746 T T T | |||
0.3751293065833128 0.8749353467083437 0.2373114907195359 T T T | |||
0.3749413149216736 0.1228899057605569 0.2378372905376746 T T T | |||
0.3750000000000000 0.3750000000000000 0.2382228949121510 T T T | |||
0.3749413149216736 0.6271687793177693 0.2378372905376746 T T T | |||
0.6254801634753705 0.8740396730492660 0.2371083546269586 T T T | |||
0.6271687793177693 0.1228899057605569 0.2378372905376746 T T T | |||
0.6271687793177693 0.3749413149216736 0.2378372905376746 T T T | |||
0.6254801634753705 0.6254801634753705 0.2371083546269586 T T T | |||
0.7916031907512163 0.7916031907512163 0.1897987884844704 T T T | |||
0.7911585061632604 0.0418454710877017 0.1901635667924394 T T T | |||
0.7911585061632604 0.2919960227490310 0.1901635667924394 T T T | |||
0.7916031907512163 0.5417936184975600 0.1897987884844704 T T T | |||
0.0418454710877017 0.7911585061632604 0.1901635667924394 T T T | |||
0.0416666666666643 0.0416666666666643 0.1902063535974311 T T T | |||
0.0418454710877017 0.2919960227490310 0.1901635667924394 T T T | |||
0.0421389046174020 0.5414305476912955 0.1898831779310922 T T T | |||
0.2919960227490310 0.7911585061632604 0.1901635667924394 T T T | |||
0.2919960227490310 0.0418454710877017 0.1901635667924394 T T T | |||
0.2918252482458862 0.2918252482458862 0.1901996737591517 T T T | |||
0.2918252482458862 0.5413495035082204 0.1901996737591517 T T T | |||
0.5417936184975600 0.7916031907512163 0.1897987884844704 T T T | |||
0.5414305476912955 0.0421389046174020 0.1898831779310922 T T T | |||
0.5413495035082204 0.2918252482458862 0.1901996737591517 T T T | |||
0.5414305476912955 0.5414305476912955 0.1898831779310922 T T T | |||
0.7083333333333357 0.7083333333333357 0.1425506314000984 F F F | |||
0.7083333333333357 0.9583333333333357 0.1425506314000984 F F F | |||
0.7083333333333357 0.2083333333333357 0.1425506314000984 F F F | |||
0.7083333333333357 0.4583333333333357 0.1425506314000984 F F F | |||
0.9583333333333357 0.7083333333333357 0.1425506314000984 F F F | |||
0.9583333333333357 0.9583333333333357 0.1425506314000984 F F F | |||
0.9583333333333357 0.2083333333333357 0.1425506314000984 F F F | |||
0.9583333333333357 0.4583333333333357 0.1425506314000984 F F F | |||
0.2083333333333357 0.7083333333333357 0.1425506314000984 F F F | |||
0.2083333333333357 0.9583333333333357 0.1425506314000984 F F F | |||
0.2083333333333357 0.2083333333333357 0.1425506314000984 F F F | |||
0.2083333333333357 0.4583333333333357 0.1425506314000984 F F F | |||
0.4583333333333357 0.7083333333333357 0.1425506314000984 F F F | |||
0.4583333333333357 0.9583333333333357 0.1425506314000984 F F F | |||
0.4583333333333357 0.2083333333333357 0.1425506314000984 F F F | |||
0.4583333333333357 0.4583333333333357 0.1425506314000984 F F F | |||
0.6250000000000000 0.6250000000000000 0.0950337542667299 F F F | |||
0.6250000000000000 0.8750000000000000 0.0950337542667299 F F F | |||
0.6250000000000000 0.1250000000000000 0.0950337542667299 F F F | |||
0.6250000000000000 0.3750000000000000 0.0950337542667299 F F F | |||
0.8750000000000000 0.6250000000000000 0.0950337542667299 F F F | |||
0.8750000000000000 0.8750000000000000 0.0950337542667299 F F F | |||
0.8750000000000000 0.1250000000000000 0.0950337542667299 F F F | |||
0.8750000000000000 0.3750000000000000 0.0950337542667299 F F F | |||
0.1250000000000000 0.6250000000000000 0.0950337542667299 F F F | |||
0.1250000000000000 0.8750000000000000 0.0950337542667299 F F F | |||
0.1250000000000000 0.1250000000000000 0.0950337542667299 F F F | |||
0.1250000000000000 0.3750000000000000 0.0950337542667299 F F F | |||
0.3750000000000000 0.6250000000000000 0.0950337542667299 F F F | |||
0.3750000000000000 0.8750000000000000 0.0950337542667299 F F F | |||
0.3750000000000000 0.1250000000000000 0.0950337542667299 F F F | |||
0.3750000000000000 0.3750000000000000 0.0950337542667299 F F F | |||
0.5416666666666643 0.5416666666666643 0.0475168771333685 F F F | |||
0.5416666666666643 0.7916666666666643 0.0475168771333685 F F F | |||
0.5416666666666643 0.0416666666666643 0.0475168771333685 F F F | |||
0.5416666666666643 0.2916666666666643 0.0475168771333685 F F F | |||
0.7916666666666643 0.5416666666666643 0.0475168771333685 F F F | |||
0.7916666666666643 0.7916666666666643 0.0475168771333685 F F F | |||
0.7916666666666643 0.0416666666666643 0.0475168771333685 F F F | |||
0.7916666666666643 0.2916666666666643 0.0475168771333685 F F F | |||
0.0416666666666643 0.5416666666666643 0.0475168771333685 F F F | |||
0.0416666666666643 0.7916666666666643 0.0475168771333685 F F F | |||
0.0416666666666643 0.0416666666666643 0.0475168771333685 F F F | |||
0.0416666666666643 0.2916666666666643 0.0475168771333685 F F F | |||
0.2916666666666643 0.5416666666666643 0.0475168771333685 F F F | |||
0.2916666666666643 0.7916666666666643 0.0475168771333685 F F F | |||
0.2916666666666643 0.0416666666666643 0.0475168771333685 F F F | |||
0.2916666666666643 0.2916666666666643 0.0475168771333685 F F F | |||
0.4583333333333357 0.4583333333333357 0.0000000000000000 F F F | |||
0.4583333333333357 0.7083333333333357 0.0000000000000000 F F F | |||
0.4583333333333357 0.9583333333333357 0.0000000000000000 F F F | |||
0.4583333333333357 0.2083333333333357 0.0000000000000000 F F F | |||
0.7083333333333357 0.4583333333333357 0.0000000000000000 F F F | |||
0.7083333333333357 0.7083333333333357 0.0000000000000000 F F F | |||
0.7083333333333357 0.9583333333333357 0.0000000000000000 F F F | |||
0.7083333333333357 0.2083333333333357 0.0000000000000000 F F F | |||
0.9583333333333357 0.4583333333333357 0.0000000000000000 F F F | |||
0.9583333333333357 0.7083333333333357 0.0000000000000000 F F F | |||
0.9583333333333357 0.9583333333333357 0.0000000000000000 F F F | |||
0.9583333333333357 0.2083333333333357 0.0000000000000000 F F F | |||
0.2083333333333357 0.4583333333333357 0.0000000000000000 F F F | |||
0.2083333333333357 0.7083333333333357 0.0000000000000000 F F F | |||
0.2083333333333357 0.9583333333333357 0.0000000000000000 F F F | |||
0.2083333333333357 0.2083333333333357 0.0000000000000000 F F F | |||
0.3750000000000000 0.3750000000000000 0.9524831228666315 F F F | |||
0.3750000000000000 0.6250000000000000 0.9524831228666315 F F F | |||
0.3750000000000000 0.8750000000000000 0.9524831228666315 F F F | |||
0.3750000000000000 0.1250000000000000 0.9524831228666315 F F F | |||
0.6250000000000000 0.3750000000000000 0.9524831228666315 F F F | |||
0.6250000000000000 0.6250000000000000 0.9524831228666315 F F F | |||
0.6250000000000000 0.8750000000000000 0.9524831228666315 F F F | |||
0.6250000000000000 0.1250000000000000 0.9524831228666315 F F F | |||
0.8750000000000000 0.3750000000000000 0.9524831228666315 F F F | |||
0.8750000000000000 0.6250000000000000 0.9524831228666315 F F F | |||
0.8750000000000000 0.8750000000000000 0.9524831228666315 F F F | |||
0.8750000000000000 0.1250000000000000 0.9524831228666315 F F F | |||
0.1250000000000000 0.3750000000000000 0.9524831228666315 F F F | |||
0.1250000000000000 0.6250000000000000 0.9524831228666315 F F F | |||
0.1250000000000000 0.8750000000000000 0.9524831228666315 F F F | |||
0.1250000000000000 0.1250000000000000 0.9524831228666315 F F F | |||
0.2916666666666643 0.2916666666666643 0.9049662457332701 F F F | |||
0.2916666666666643 0.5416666666666643 0.9049662457332701 F F F | |||
0.2916666666666643 0.7916666666666643 0.9049662457332701 F F F | |||
0.2916666666666643 0.0416666666666643 0.9049662457332701 F F F | |||
0.5416666666666643 0.2916666666666643 0.9049662457332701 F F F | |||
0.5416666666666643 0.5416666666666643 0.9049662457332701 F F F | |||
0.5416666666666643 0.7916666666666643 0.9049662457332701 F F F | |||
0.5416666666666643 0.0416666666666643 0.9049662457332701 F F F | |||
0.7916666666666643 0.2916666666666643 0.9049662457332701 F F F | |||
0.7916666666666643 0.5416666666666643 0.9049662457332701 F F F | |||
0.7916666666666643 0.7916666666666643 0.9049662457332701 F F F | |||
0.7916666666666643 0.0416666666666643 0.9049662457332701 F F F | |||
0.0416666666666643 0.2916666666666643 0.9049662457332701 F F F | |||
0.0416666666666643 0.5416666666666643 0.9049662457332701 F F F | |||
0.0416666666666643 0.7916666666666643 0.9049662457332701 F F F | |||
0.0416666666666643 0.0416666666666643 0.9049662457332701 F F F | |||
0.2083333333333357 0.2083333333333357 0.8574493685999016 F F F | |||
0.2083333333333357 0.4583333333333357 0.8574493685999016 F F F | |||
0.2083333333333357 0.7083333333333357 0.8574493685999016 F F F | |||
0.2083333333333357 0.9583333333333357 0.8574493685999016 F F F | |||
0.4583333333333357 0.2083333333333357 0.8574493685999016 F F F | |||
0.4583333333333357 0.4583333333333357 0.8574493685999016 F F F | |||
0.4583333333333357 0.7083333333333357 0.8574493685999016 F F F | |||
0.4583333333333286 0.9583333333333357 0.8574493685999016 F F F | |||
0.7083333333333357 0.2083333333333357 0.8574493685999016 F F F | |||
0.7083333333333357 0.4583333333333357 0.8574493685999016 F F F | |||
0.7083333333333357 0.7083333333333357 0.8574493685999016 F F F | |||
0.7083333333333286 0.9583333333333357 0.8574493685999016 F F F | |||
0.9583333333333357 0.2083333333333357 0.8574493685999016 F F F | |||
0.9583333333333357 0.4583333333333357 0.8574493685999016 F F F | |||
0.9583333333333357 0.7083333333333357 0.8574493685999016 F F F | |||
0.9583333333333357 0.9583333333333357 0.8574493685999016 F F F | |||
0.1248753274088950 0.1248753274088950 0.8104723183369446 T T T | |||
0.1250432471286117 0.3744669037981175 0.8104439099895814 T T T | |||
0.1250432471286117 0.6254898490732638 0.8104439099895814 T T T | |||
0.1248753274088950 0.8752493451822171 0.8104723183369446 T T T | |||
0.3744669037981175 0.1250432471286117 0.8104439099895814 T T T | |||
0.3750000000000000 0.3750000000000000 0.8101623576913966 T T T | |||
0.3744669037981175 0.6254898490732638 0.8104439099895814 T T T | |||
0.3741437503952032 0.8754281248023950 0.8106074679484812 T T T | |||
0.6254898490732638 0.1250432471286117 0.8104439099895814 T T T | |||
0.6254898490732638 0.3744669037981175 0.8104439099895814 T T T | |||
0.6249056987982424 0.6249056987982424 0.8109175909471307 T T T | |||
0.6249056987982424 0.8751886024035143 0.8109175909471307 T T T | |||
0.8752493451822171 0.1248753274088950 0.8104723183369446 T T T | |||
0.8754281248023950 0.3741437503952032 0.8106074679484812 T T T | |||
0.8751886024035143 0.6249056987982424 0.8109175909471307 T T T | |||
0.8754281248023950 0.8754281248023950 0.8106074679484812 T T T | |||
0.0416666666666643 0.0416666666666643 0.7630725963525371 T T T | |||
0.0423582548037352 0.2929829466331252 0.7632255425190596 T T T | |||
0.0423342079934537 0.5413328960032665 0.7639551649056903 T T T | |||
0.0423582548037352 0.7896587985631328 0.7632255425190596 T T T | |||
0.2929829466331252 0.0423582548037352 0.7632255425190596 T T T | |||
0.2921448802009530 0.2921448802009530 0.7631719676853072 T T T | |||
0.2921448802009530 0.5407102395980798 0.7631719676853072 T T T | |||
0.2929829466331252 0.7896587985631328 0.7632255425190596 T T T | |||
0.5413328960032665 0.0423342079934537 0.7639551649056903 T T T | |||
0.5407102395980798 0.2921448802009530 0.7631719676853072 T T T | |||
0.5413328960032665 0.5413328960032665 0.7639551649056903 T T T | |||
0.5423722548894060 0.7913138725552938 0.7643637592766812 T T T | |||
0.7896587985631328 0.0423582548037352 0.7632255425190596 T T T | |||
0.7896587985631328 0.2929829466331252 0.7632255425190596 T T T | |||
0.7913138725552938 0.5423722548894060 0.7643637592766812 T T T | |||
0.7913138725552938 0.7913138725552938 0.7643637592766812 T T T | |||
0.9592783288347152 0.9592783288347152 0.7165014402517978 T T T | |||
0.9592783288347081 0.2064433423305768 0.7165014402517978 T T T | |||
0.9585429811439916 0.4580852035394961 0.7171342391736614 T T T | |||
0.9585429811439987 0.7083718153165048 0.7171342391736614 T T T | |||
0.2064433423305768 0.9592783288347152 0.7165014402517978 T T T | |||
0.2080072672259897 0.2080072672259968 0.7154393564109170 T T T | |||
0.2098510312054594 0.4575744843972738 0.7163450633092348 T T T | |||
0.2080072672259968 0.7089854655480138 0.7154393564109170 T T T | |||
0.4580852035394889 0.9585429811439987 0.7171342391736614 T T T | |||
0.4575744843972738 0.2098510312054594 0.7163450633092348 T T T | |||
0.4575744843972738 0.4575744843972738 0.7163450633092348 T T T | |||
0.4580852035394961 0.7083718153165048 0.7171342391736614 T T T | |||
0.7083718153165048 0.9585429811439987 0.7171342391736614 T T T | |||
0.7089854655480138 0.2080072672259968 0.7154393564109170 T T T | |||
0.7083718153165048 0.4580852035394961 0.7171342391736614 T T T | |||
0.7083333333333357 0.7083333333333357 0.7173121617349102 T T T | |||
0.9312769255399772 0.9312769255399772 0.3485062381768784 T T T | |||
0.9312769255399772 0.2624461489200455 0.3485062381768784 T T T | |||
0.9325931874642169 0.5962034062678949 0.3492759583004709 T T T | |||
0.2624461489200455 0.9312769255399772 0.3485062381768784 T T T | |||
0.2649012212242361 0.2649012212242361 0.3482925057539119 T T T | |||
0.2649012212242361 0.5951975575515207 0.3482925057539119 T T T | |||
0.5962034062678949 0.9325931874642169 0.3492759583004709 T T T | |||
0.5951975575515207 0.2649012212242361 0.3482925057539119 T T T | |||
0.5962034062678949 0.5962034062678949 0.3492759583004709 T T T | |||
0.8194444444444429 0.8194444444444429 0.4120073015393046 F F F | |||
0.8194444444444429 0.1527777777777786 0.4120073015393046 F F F | |||
0.8194444444444429 0.4861111111111143 0.4120073015393046 F F F | |||
0.1527777777777786 0.8194444444444429 0.4120073015393046 F F F | |||
0.1527777777777786 0.1527777777777786 0.4120073015393046 F F F | |||
0.1527777777777786 0.4861111111111143 0.4120073015393046 F F F | |||
0.4861111111111143 0.8194444444444429 0.4120073015393046 F F F | |||
0.4861111111111143 0.1527777777777786 0.4120073015393046 F F F | |||
0.4861111111111143 0.4861111111111143 0.4120073015393046 F F F | |||
0.7083333333333357 0.7083333333333357 0.4760019913289000 F F F | |||
0.7083333333333357 0.0416666666666643 0.4760019913289000 F F F | |||
0.7083333333333357 0.3750000000000000 0.4760019913289000 F F F | |||
0.0416666666666643 0.7083333333333357 0.4760019913289000 F F F | |||
0.0416666666666643 0.0416666666666643 0.4760019913289000 F F F | |||
0.0416666666666643 0.3750000000000000 0.4760019913289000 F F F | |||
0.3750000000000000 0.7083333333333357 0.4760019913289000 F F F | |||
0.3750000000000000 0.0416666666666643 0.4760019913289000 F F F | |||
0.3750000000000000 0.3750000000000000 0.4760019913289000 F F F | |||
0.5972222222222214 0.5972222222222214 0.5399966811184953 F F F | |||
0.5972222222222214 0.9305555555555571 0.5399966811184953 F F F | |||
0.5972222222222214 0.2638888888888857 0.5399966811184953 F F F | |||
0.9305555555555571 0.5972222222222214 0.5399966811184953 F F F | |||
0.9305555555555571 0.9305555555555571 0.5399966811184953 F F F | |||
0.9305555555555571 0.2638888888888857 0.5399966811184953 F F F | |||
0.2638888888888857 0.5972222222222214 0.5399966811184953 F F F | |||
0.2638888888888857 0.9305555555555571 0.5399966811184953 F F F | |||
0.2638888888888857 0.2638888888888857 0.5399966811184953 F F F | |||
0.4861111111111143 0.4861111111111143 0.6039913709080977 F F F | |||
0.4861111111111143 0.8194444444444429 0.6039913709080977 F F F | |||
0.4861111111111143 0.1527777777777786 0.6039913709080977 F F F | |||
0.8194444444444429 0.4861111111111143 0.6039913709080977 F F F | |||
0.8194444444444429 0.8194444444444429 0.6039913709080977 F F F | |||
0.8194444444444429 0.1527777777777786 0.6039913709080977 F F F | |||
0.1527777777777786 0.4861111111111143 0.6039913709080977 F F F | |||
0.1527777777777786 0.8194444444444429 0.6039913709080977 F F F | |||
0.1527777777777786 0.1527777777777786 0.6039913709080977 F F F | |||
0.3750000000000000 0.3750000000000000 0.6709451889760421 T T T | |||
0.3757686837881531 0.7078847317116083 0.6692728159340477 T T T | |||
0.3757686837881531 0.0413465845002455 0.6692728159340477 T T T | |||
0.7078847317116083 0.3757686837881531 0.6692728159340477 T T T | |||
0.7083333333333357 0.7083333333333357 0.6672815507555968 T T T | |||
0.7078847317116083 0.0413465845002455 0.6692728159340477 T T T | |||
0.0413465845002455 0.3757686837881531 0.6692728159340477 T T T | |||
0.0413465845002455 0.7078847317116083 0.6692728159340477 T T T | |||
0.0416666666666643 0.0416666666666643 0.6705979807351840 T T T | |||
0.0416666666666643 0.0416666666666643 0.3295224610717508 T T T | |||
0.0415965476045645 0.3744525699724842 0.3318913105762326 T T T | |||
0.0415965476045645 0.7089508824229516 0.3318913105762326 T T T | |||
0.3744525699724842 0.0415965476045645 0.3318913105762326 T T T | |||
0.3750000000000000 0.3750000000000000 0.3298742237993510 T T T | |||
0.3744525699724842 0.7089508824229516 0.3318913105762326 T T T | |||
0.7089508824229516 0.0415965476045645 0.3318913105762326 T T T | |||
0.7089508824229516 0.3744525699724842 0.3318913105762326 T T T | |||
0.7083333333333357 0.7083333333333357 0.3343592162472642 T T T | |||
0.9305555555555571 0.9305555555555571 0.3960086290919023 F F F | |||
0.9305555555555571 0.2638888888888857 0.3960086290919023 F F F | |||
0.9305555555555571 0.5972222222222214 0.3960086290919023 F F F | |||
0.2638888888888857 0.9305555555555571 0.3960086290919023 F F F | |||
0.2638888888888857 0.2638888888888857 0.3960086290919023 F F F | |||
0.2638888888888857 0.5972222222222214 0.3960086290919023 F F F | |||
0.5972222222222214 0.9305555555555571 0.3960086290919023 F F F | |||
0.5972222222222214 0.2638888888888857 0.3960086290919023 F F F | |||
0.5972222222222214 0.5972222222222214 0.3960086290919023 F F F | |||
0.8194444444444429 0.8194444444444429 0.4600033188815047 F F F | |||
0.8194444444444429 0.1527777777777786 0.4600033188815047 F F F | |||
0.8194444444444429 0.4861111111111143 0.4600033188815047 F F F | |||
0.1527777777777786 0.8194444444444429 0.4600033188815047 F F F | |||
0.1527777777777786 0.1527777777777786 0.4600033188815047 F F F | |||
0.1527777777777786 0.4861111111111143 0.4600033188815047 F F F | |||
0.4861111111111143 0.8194444444444429 0.4600033188815047 F F F | |||
0.4861111111111143 0.1527777777777786 0.4600033188815047 F F F | |||
0.4861111111111143 0.4861111111111143 0.4600033188815047 F F F | |||
0.7083333333333357 0.7083333333333357 0.5239980086711000 F F F | |||
0.7083333333333357 0.0416666666666643 0.5239980086711000 F F F | |||
0.7083333333333357 0.3750000000000000 0.5239980086711000 F F F | |||
0.0416666666666643 0.7083333333333357 0.5239980086711000 F F F | |||
0.0416666666666643 0.0416666666666643 0.5239980086711000 F F F | |||
0.0416666666666643 0.3750000000000000 0.5239980086711000 F F F | |||
0.3750000000000000 0.7083333333333357 0.5239980086711000 F F F | |||
0.3750000000000000 0.0416666666666643 0.5239980086711000 F F F | |||
0.3750000000000000 0.3750000000000000 0.5239980086711000 F F F | |||
0.5972222222222214 0.5972222222222214 0.5879926984606954 F F F | |||
0.5972222222222214 0.9305555555555571 0.5879926984606954 F F F | |||
0.5972222222222214 0.2638888888888857 0.5879926984606954 F F F | |||
0.9305555555555571 0.5972222222222214 0.5879926984606954 F F F | |||
0.9305555555555571 0.9305555555555571 0.5879926984606954 F F F | |||
0.9305555555555571 0.2638888888888857 0.5879926984606954 F F F | |||
0.2638888888888857 0.5972222222222214 0.5879926984606954 F F F | |||
0.2638888888888857 0.9305555555555571 0.5879926984606954 F F F | |||
0.2638888888888857 0.2638888888888857 0.5879926984606954 F F F | |||
0.4857962518454938 0.4857962518454938 0.6520134262762931 T T T | |||
0.4849632646245893 0.8200183676877012 0.6512691959389023 T T T | |||
0.4857962518454938 0.1534074963090196 0.6520134262762931 T T T | |||
0.8200183676877012 0.4849632646245893 0.6512691959389023 T T T | |||
0.8200183676877012 0.8200183676877012 0.6512691959389023 T T T | |||
0.8206112061993666 0.1521943969003203 0.6522258679919717 T T T | |||
0.1534074963090195 0.4857962518454938 0.6520134262762931 T T T | |||
0.1521943969003203 0.8206112061993666 0.6522258679919717 T T T | |||
0.1521943969003203 0.1521943969003203 0.6522258679919717 T T T | |||
</pre> | |||
</div> | |||
After computing the electrostatic potential by setting {{TAG|WRT_POTENTIAL|hartree ionic}}, {{TAG|PREC|Accurate}}, and {{TAG|KSPACING|0.2}}, we can run the Python script referenced above in Step 4. | |||
For the Al(111)/Si(111) interface studied, we get the following results: <math>\bar{V}_{\mathrm{m}} = -0.694</math> eV, <math>\bar{V}_{\mathrm{sc}} = +1.192</math> eV, and <math>\Delta\bar{V} = -1.886</math> eV. | |||
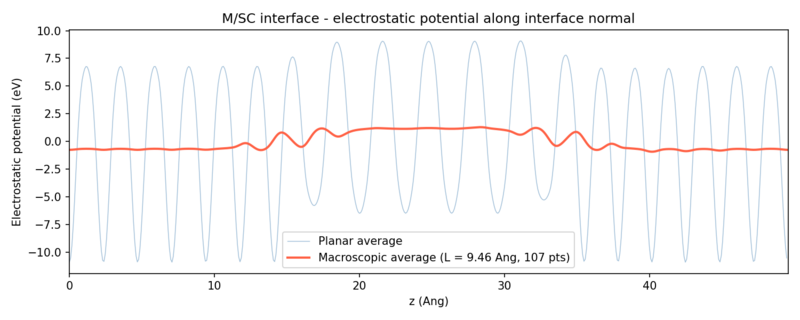
[[File:Potential alignment Si111 Al111.png|thumbnail|800px|center|Potential alignment plot for the Al(111) Si(111) interface.]] | |||
Finally, we arrive at the Schottky barriers: | |||
:<math>\varphi_{\mathrm{p}} = \Delta\bar{V} + E_{\mathrm{F}} - E_{\mathrm{VBM}} = -1.886 + 8.085 - 5.449 = 0.75</math> eV, | |||
which fits well to the experimental values that vary between 0.68 and 0.81 eV, depending on sample and experimental conditions{{cite|altindal:2007}}, and | |||
:<math>\varphi_{\mathrm{n}} = E_{\mathrm{g}} - \varphi_{\mathrm{p}} = 1.160 - 0.75 = 0.41</math> eV. | |||
== Related tags and articles == | == Related tags and articles == | ||
{{TAG|LVHAR}}, | |||
{{TAG|WRT_POTENTIAL}}, | |||
{{TAG|BANDGAP}}, | |||
[[Computing the work function]] | |||
== References == | == References == | ||
[[Category:Howto]] | |||
Latest revision as of 12:22, 27 March 2026
The Schottky barrier is the energy barrier that forms at a metal–semiconductor junction. The barrier height is an important characteristic for charge transport across the junction and a critical parameter in the design of semiconductor contacts, transistors, and rectifying diodes. The p-type Schottky barrier height [math]\displaystyle{ \varphi_{\mathrm{p}} }[/math] describes the barrier seen by holes in the semiconductor valence band, and the n-type Schottky barrier height [math]\displaystyle{ \varphi_{\mathrm{n}} }[/math] describes the barrier seen by electrons in the conduction band. In VASP, the potential alignment method can be used to estimate the Schottky barrier[1][2].
Required quantities
Three quantities from bulk calculations, the Fermi energy [math]\displaystyle{ E_{\mathrm{F}} }[/math] of the metal, and the valence-band maximum [math]\displaystyle{ E_{\mathrm{VBM}} }[/math] and band gap [math]\displaystyle{ E_{\mathrm{g}} }[/math] of the semiconductor, are needed for the calculation. In addition, the macroscopic electrostatic potential difference [math]\displaystyle{ \Delta\bar{V} }[/math] across the interface is extracted from a separate interface slab calculation[3].
Step-by-step instructions
| Important: For the calculation of the p- and n-type Schottky barriers, energies, and potentials from different VASP calculations are compared. It is crucial to set a single value for ENCUT for all calculations. |
Step 1: Compute the Fermi energy of the metal.
- After ensuring that the bulk geometry is optimized, a separate static calculation should be performed with a dense, Γ-centered k-point mesh and
ISMEAR = -5(tetrahedron method with Blöchl corrections), to get an accurate Fermi energy. It can be taken from the OUTCAR and is also easily accessible via py4vasp: import py4vasp as pv calc = pv.Calculation.from_path("./Step1/") E_F = calc.dos.read()["fermi_energy"] print(f"E_F = {E_F:.4f} eV")
Step 2: Compute the band gap and valence-band maximum of the semiconductor.
- Band gaps are notoriously underestimated by standard DFT, so for accurate values, it is usually necessary to use DFT+U, hybrid functionals, or GW calculations. The level of theory to choose depends on the semiconductor of interest, and HSE06 is oftern a good default for most covalent and ionic semiconductors.
- Generally, it is advisable to select
BANDGAP = KPOINTto get more verbose output, and ideally to make a full band-structure calculation, since the extrema of the bands are often located at high-symmetry lines. The fundamental bandgap [math]\displaystyle{ E_{\mathrm{g}} }[/math] and the valence-band maximum (VBM) [math]\displaystyle{ E_{\mathrm{VBM}} }[/math], can be read from the OUTCAR file, or with py4vasp: import py4vasp as pv calc = pv.Calculation.from_path("./Step2/") E_g = calc.bandgap.fundamental() E_VBM = calc.bandgap.valence_band_maximum() print(f"E_g = {E_g:.4f} eV") print(f"E_VBM = {E_VBM:.4f} eV")
Step 3: Build and relax an interface structure.
- Building a lattice-matched metal–semiconductor interface slab is outside the scope of this page. Tools such as the CoherentInterfaceBuilder in pymatgen[4] and the interface construction utilities in ASE[5] can be used to construct the initial geometry. An interface structure for a Schottky barrier calculation should satisfy the following requirements:
- Low lattice mismatch. The supercell should be chosen such that the in-plane periodicities of the two surfaces match to within ~2%, minimising artificial biaxial strain.
- Sufficient slab thickness. Each material should contain enough layers that a bulk-like region exists in the centre.
- No vacuum region. A cell with a vacuum will result in lateral strain, compressing the cell to reduce the surface energy. It is better to have two equivalent interfaces in the cell without any vacuum at all.
- For the relaxation, it is usually prudent to fix the layers in the bulk like regions using selective dynamics in the POSCAR file. Only a couple of layers at the interfaces should be relaxed. Depending on the material pairing and the strain in the interface, it might be beneficial to allow the cell to relax only in the direction normal to the interface plane using the LATTICE_CONSTRAINTS tag or the ICONST file to apply the contraint.
Step 4: Compute the electrostatic potential of the interface and calculate the Schottky barrier heights.
- Perform a static calculation with tetrahedron smearing (
ISMEAR = -5) andPREC = Accurate. We need to print out the electrostatic part of the potential (disregarding the XC part) by settingWRT_POTENTIAL = hartree ionic. It is also possible to useLVHAR = .TRUE.to get the same data, but be aware that LVTOT would include the (semi-)local exchange-correlation potential [math]\displaystyle{ V_{\text{xc}}(\mathbf{r}) }[/math], which we want to exclude here.
- The planar average [math]\displaystyle{ \bar{V}(z) }[/math] of the electrostatic potential is obtained by averaging the 3D potential over the in-plane (xy) grid at each [math]\displaystyle{ z }[/math] point (assuming the interface is located in the xy-plane). Then, a macroscopic average is computed by applying a uniform box-car convolution of window length [math]\displaystyle{ L }[/math], where [math]\displaystyle{ L }[/math] equals one bulk primitive-cell repeat period along the interface normal. Averaging over exactly one such period removes the short-range oscillations within each atomic layer while preserving the long-range potential step across the interface. The Python script below attempts to find the optimal [math]\displaystyle{ L }[/math] automatically by sweeping a range of values and selecting the one that minimizes the RMS gradient of the macroscopic average in the bulk-like plateau regions of both materials simultaneously. It reads the potential from vaspout.h5, computes both averages, plots the data and reports the macroscopic electrostatic potential difference [math]\displaystyle{ \Delta\bar{V} }[/math]:
- Click to show python script
#!/usr/bin/env python3 """ Potential alignment script for Schottky barrier height calculations. The electrostatic potential (hartree + ionic) is read from vaspout.h5, averaged in the xy-plane to give the planar average V_bar(z), and then smoothed with a box-car macroscopic average of width L to remove short-range oscillations. dV_bar = <V_bar>_M - <V_bar>_SC is the potential offset between the metal (M) and semiconductor (SC) bulk-like regions of the slab. L is chosen automatically by minimising the worst-case RMS gradient in the two plateau regions (M and SC separately). This ensures both regions are flat, even when the two materials have different lattice periodicities along z. The plateau centres are given as fractional cell coordinates via --mc (metal) and --scc (semiconductor). A fixed plateau half-width of 0.08 * c is used. """ import argparse import matplotlib matplotlib.use("Agg") # non-interactive backend; remove if you want a GUI window import matplotlib.pyplot as plt import h5py import numpy as np from scipy.ndimage import uniform_filter1d parser = argparse.ArgumentParser(description="Compute macroscopic potential alignment.") parser.add_argument( "--mc", "--metal-center", type=float, default=0.125, metavar="F", dest="metal_center", help="Fractional z-coordinate of the metal plateau centre (default: 0.125)", ) parser.add_argument( "--scc", "--sc-center", type=float, default=0.5, metavar="F", dest="sc_center", help="Fractional z-coordinate of the semiconductor plateau centre (default: 0.5)", ) parser.add_argument( "--vaspout", default="./Step4/vaspout.h5", metavar="FILE", help="Path to vaspout.h5 (default: ./vaspout.h5)", ) args = parser.parse_args() # -- load potential and cell geometry ------------------------------------------ with h5py.File(args.vaspout, "r") as f: hartree = f["/results/potential/hartree"][0] # (NGZ, NGY, NGX) ionic = f["/results/potential/ionic"][0] # (NGZ, NGY, NGX) lat = f["/results/positions/lattice_vectors"][:] # (3, 3) Ang c = np.linalg.norm(lat[2]) # cell length along interface normal (Ang) NGZ = hartree.shape[0] # number of grid points along z dz = c / NGZ # grid spacing (Ang) # -- planar average: mean over the xy-plane at each z -------------------------- V_planar = (hartree + ionic).mean(axis=(1, 2)) # shape (NGZ,) [eV] z = np.arange(NGZ) * dz # z-coordinates in Ang # -- plateau masks (used both for auto-selection and for dV_bar extraction) ---- # z_frac maps each grid point to a fractional cell coordinate in [0, 1). # For each material the mask selects a symmetric window of width 2*HALF_WIDTH # centred on the user-supplied fractional coordinate (--mc / --scc). The # distance is computed with periodic boundary conditions (minimum image), so # --mc=0.0 correctly selects both the bottom edge (z_frac < 0.08) and the # top edge (z_frac > 0.92) of the cell, which belong to the same metal region. # HALF_WIDTH = 0.08 means the window covers 16 % of the cell on each side, # which is large enough to average over several bulk-like repeat units while # staying well clear of the interface region for any reasonably converged # slab (total cell >= ~40 Ang, >= 6 layers per material). HALF_WIDTH = 0.08 z_frac = z / c def _periodic_mask(center): d = np.abs(z_frac - center) d = np.minimum(d, 1.0 - d) # minimum-image distance in fractional coords return d < HALF_WIDTH M_mask = _periodic_mask(args.metal_center) SC_mask = _periodic_mask(args.sc_center) # -- auto-select window length ------------------------------------------------- # Sweep L from 2*dz to c/3. For each L the score is the worst-case RMS gradient # across the two plateau regions (M and SC scored independently). Minimising the # worst case forces both regions to be flat simultaneously. L_grid = np.linspace(2 * dz, c / 3, 300) def _plateau_roughness(L): Vm = uniform_filter1d(V_planar, size=max(1, int(round(L / dz))), mode="wrap") g = np.gradient(Vm) return max(np.sqrt(np.mean(g[M_mask]**2)), np.sqrt(np.mean(g[SC_mask]**2))) scores = [_plateau_roughness(L) for L in L_grid] WINDOW_LENGTH = float(L_grid[np.argmin(scores)]) print(f"Auto-selected window length: {WINDOW_LENGTH:.2f} Ang") # -- macroscopic average: box-car convolution with period L -------------------- # uniform_filter1d computes the running mean over `size` consecutive points. # mode='wrap' enforces periodic boundary conditions, which is correct for # a slab calculation where the potential is periodic along z. n_window = int(round(WINDOW_LENGTH / dz)) V_macro = uniform_filter1d(V_planar, size=n_window, mode="wrap") # -- plot ---------------------------------------------------------------------- fig, ax = plt.subplots(figsize=(10, 4)) ax.plot(z, V_planar, lw=0.7, color="steelblue", alpha=0.5, label="Planar average") ax.plot(z, V_macro, lw=2.0, color="tomato", label=f"Macroscopic average (L = {WINDOW_LENGTH:.2f} Ang, {n_window} pts)") ax.set_xlabel("z (Ang)") ax.set_ylabel("Electrostatic potential (eV)") ax.set_xlim(0, c) ax.legend() ax.set_title("M/SC interface - electrostatic potential along interface normal") plt.tight_layout() plt.savefig("potential_alignment.png", dpi=150) print("Figure saved: potential_alignment.png") # -- extract dV_bar from bulk-like plateaus ------------------------------------ # M_mask and SC_mask are defined above; adjust --mc / --scc if the plateaus shift. V_M = V_macro[M_mask].mean() V_SC = V_macro[SC_mask].mean() delta_V = V_M - V_SC print(f"\n<V_bar>_M = {V_M:+.4f} eV") print(f"<V_bar>_SC = {V_SC:+.4f} eV") print(f"dV_bar = <V_bar>_M - <V_bar>_SC = {delta_V:+.4f} eV")
Step 5: Compute the Schottky barrier heights.
- With [math]\displaystyle{ E_{\mathrm{F}} }[/math], [math]\displaystyle{ E_{\mathrm{VBM}} }[/math], [math]\displaystyle{ E_{\mathrm{g}} }[/math], and [math]\displaystyle{ \Delta\bar{V} }[/math] in hand, the potential alignment difference [math]\displaystyle{ \Delta\bar{V} = \bar{V}_{\mathrm{m}} - \bar{V}_{\mathrm{sc}} }[/math] connects the bulk reference frames. The p-type and n-type Schottky barrier heights are then
- [math]\displaystyle{ \varphi_{\mathrm{p}} = \Delta\bar{V} + E_{\mathrm{F}} - E_{\mathrm{VBM}}, }[/math]
- [math]\displaystyle{ \varphi_{\mathrm{n}} = E_{\mathrm{g}} - \varphi_{\mathrm{p}}. }[/math]
Example
Here we will consider the Al(111)/Si(111) interface. The bulk Al calculation to determine the Fermi energy is straightforward and yields [math]\displaystyle{ E_{\mathrm{F}}=8.085 }[/math] eV for the PBE functional with ENCUT = 300, KSPACING = 0.2, and ISMEAR = -5.
For the Si bandgap and valence band maximum, a little more care is needed. After relaxing with PBEsol to get the correct lattice parameter, a band structure calculation with the HSE06 functional yields a bandgap of [math]\displaystyle{ E_{\mathrm{g}}=1.160 }[/math] eV and a valence band maximum of [math]\displaystyle{ E_{\mathrm{VBM}}=5.449 }[/math] eV.
For Al(111) and Si(111), a 4×4 Al supercell matched to a 3×3 Si supercell achieves a mismatch of 0.9%. For this example, we chose 13 Al layers, and 6 Si bilayers. The starting interface distance before relaxation was set to 2.3 Å We have fully relaxed the cell, but fixed all but the 4 layers closest to the interfaces (2 Al layers, and 1 Si bi-layer for each interface) with the selective dynamics POSCAR option. The relaxed structure with its 316 atoms is given here:
Al208 Si108
1.00000000000000
11.4573925481650338 0.0000000000000000 0.0000000000000007
5.7286962740825134 9.9223930078414337 0.0000000000000007
-0.0000000000000000 -0.0000000000000000 49.5333046267364523
Al Si
208 108
Selective dynamics
Direct
0.9583333333333357 0.9583333333333357 0.2851012628001968 F F F
0.9583333333333357 0.2083333333333357 0.2851012628001968 F F F
0.9583333333333357 0.4583333333333357 0.2851012628001968 F F F
0.9583333333333357 0.7083333333333357 0.2851012628001968 F F F
0.2083333333333357 0.9583333333333357 0.2851012628001968 F F F
0.2083333333333357 0.2083333333333357 0.2851012628001968 F F F
0.2083333333333357 0.4583333333333357 0.2851012628001968 F F F
0.2083333333333357 0.7083333333333357 0.2851012628001968 F F F
0.4583333333333357 0.9583333333333357 0.2851012628001968 F F F
0.4583333333333357 0.2083333333333357 0.2851012628001968 F F F
0.4583333333333357 0.4583333333333357 0.2851012628001968 F F F
0.4583333333333357 0.7083333333333357 0.2851012628001968 F F F
0.7083333333333357 0.9583333333333357 0.2851012628001968 F F F
0.7083333333333357 0.2083333333333357 0.2851012628001968 F F F
0.7083333333333357 0.4583333333333357 0.2851012628001968 F F F
0.7083333333333357 0.7083333333333357 0.2851012628001968 F F F
0.8749353467083437 0.8749353467083437 0.2373114907195359 T T T
0.8763358121166870 0.1243320939416563 0.2376670183932849 T T T
0.8749353467083437 0.3751293065833128 0.2373114907195359 T T T
0.8740396730492660 0.6254801634753705 0.2371083546269586 T T T
0.1243320939416563 0.8763358121166870 0.2376670183932849 T T T
0.1243320939416563 0.1243320939416563 0.2376670183932849 T T T
0.1228899057605569 0.3749413149216736 0.2378372905376746 T T T
0.1228899057605569 0.6271687793177693 0.2378372905376746 T T T
0.3751293065833128 0.8749353467083437 0.2373114907195359 T T T
0.3749413149216736 0.1228899057605569 0.2378372905376746 T T T
0.3750000000000000 0.3750000000000000 0.2382228949121510 T T T
0.3749413149216736 0.6271687793177693 0.2378372905376746 T T T
0.6254801634753705 0.8740396730492660 0.2371083546269586 T T T
0.6271687793177693 0.1228899057605569 0.2378372905376746 T T T
0.6271687793177693 0.3749413149216736 0.2378372905376746 T T T
0.6254801634753705 0.6254801634753705 0.2371083546269586 T T T
0.7916031907512163 0.7916031907512163 0.1897987884844704 T T T
0.7911585061632604 0.0418454710877017 0.1901635667924394 T T T
0.7911585061632604 0.2919960227490310 0.1901635667924394 T T T
0.7916031907512163 0.5417936184975600 0.1897987884844704 T T T
0.0418454710877017 0.7911585061632604 0.1901635667924394 T T T
0.0416666666666643 0.0416666666666643 0.1902063535974311 T T T
0.0418454710877017 0.2919960227490310 0.1901635667924394 T T T
0.0421389046174020 0.5414305476912955 0.1898831779310922 T T T
0.2919960227490310 0.7911585061632604 0.1901635667924394 T T T
0.2919960227490310 0.0418454710877017 0.1901635667924394 T T T
0.2918252482458862 0.2918252482458862 0.1901996737591517 T T T
0.2918252482458862 0.5413495035082204 0.1901996737591517 T T T
0.5417936184975600 0.7916031907512163 0.1897987884844704 T T T
0.5414305476912955 0.0421389046174020 0.1898831779310922 T T T
0.5413495035082204 0.2918252482458862 0.1901996737591517 T T T
0.5414305476912955 0.5414305476912955 0.1898831779310922 T T T
0.7083333333333357 0.7083333333333357 0.1425506314000984 F F F
0.7083333333333357 0.9583333333333357 0.1425506314000984 F F F
0.7083333333333357 0.2083333333333357 0.1425506314000984 F F F
0.7083333333333357 0.4583333333333357 0.1425506314000984 F F F
0.9583333333333357 0.7083333333333357 0.1425506314000984 F F F
0.9583333333333357 0.9583333333333357 0.1425506314000984 F F F
0.9583333333333357 0.2083333333333357 0.1425506314000984 F F F
0.9583333333333357 0.4583333333333357 0.1425506314000984 F F F
0.2083333333333357 0.7083333333333357 0.1425506314000984 F F F
0.2083333333333357 0.9583333333333357 0.1425506314000984 F F F
0.2083333333333357 0.2083333333333357 0.1425506314000984 F F F
0.2083333333333357 0.4583333333333357 0.1425506314000984 F F F
0.4583333333333357 0.7083333333333357 0.1425506314000984 F F F
0.4583333333333357 0.9583333333333357 0.1425506314000984 F F F
0.4583333333333357 0.2083333333333357 0.1425506314000984 F F F
0.4583333333333357 0.4583333333333357 0.1425506314000984 F F F
0.6250000000000000 0.6250000000000000 0.0950337542667299 F F F
0.6250000000000000 0.8750000000000000 0.0950337542667299 F F F
0.6250000000000000 0.1250000000000000 0.0950337542667299 F F F
0.6250000000000000 0.3750000000000000 0.0950337542667299 F F F
0.8750000000000000 0.6250000000000000 0.0950337542667299 F F F
0.8750000000000000 0.8750000000000000 0.0950337542667299 F F F
0.8750000000000000 0.1250000000000000 0.0950337542667299 F F F
0.8750000000000000 0.3750000000000000 0.0950337542667299 F F F
0.1250000000000000 0.6250000000000000 0.0950337542667299 F F F
0.1250000000000000 0.8750000000000000 0.0950337542667299 F F F
0.1250000000000000 0.1250000000000000 0.0950337542667299 F F F
0.1250000000000000 0.3750000000000000 0.0950337542667299 F F F
0.3750000000000000 0.6250000000000000 0.0950337542667299 F F F
0.3750000000000000 0.8750000000000000 0.0950337542667299 F F F
0.3750000000000000 0.1250000000000000 0.0950337542667299 F F F
0.3750000000000000 0.3750000000000000 0.0950337542667299 F F F
0.5416666666666643 0.5416666666666643 0.0475168771333685 F F F
0.5416666666666643 0.7916666666666643 0.0475168771333685 F F F
0.5416666666666643 0.0416666666666643 0.0475168771333685 F F F
0.5416666666666643 0.2916666666666643 0.0475168771333685 F F F
0.7916666666666643 0.5416666666666643 0.0475168771333685 F F F
0.7916666666666643 0.7916666666666643 0.0475168771333685 F F F
0.7916666666666643 0.0416666666666643 0.0475168771333685 F F F
0.7916666666666643 0.2916666666666643 0.0475168771333685 F F F
0.0416666666666643 0.5416666666666643 0.0475168771333685 F F F
0.0416666666666643 0.7916666666666643 0.0475168771333685 F F F
0.0416666666666643 0.0416666666666643 0.0475168771333685 F F F
0.0416666666666643 0.2916666666666643 0.0475168771333685 F F F
0.2916666666666643 0.5416666666666643 0.0475168771333685 F F F
0.2916666666666643 0.7916666666666643 0.0475168771333685 F F F
0.2916666666666643 0.0416666666666643 0.0475168771333685 F F F
0.2916666666666643 0.2916666666666643 0.0475168771333685 F F F
0.4583333333333357 0.4583333333333357 0.0000000000000000 F F F
0.4583333333333357 0.7083333333333357 0.0000000000000000 F F F
0.4583333333333357 0.9583333333333357 0.0000000000000000 F F F
0.4583333333333357 0.2083333333333357 0.0000000000000000 F F F
0.7083333333333357 0.4583333333333357 0.0000000000000000 F F F
0.7083333333333357 0.7083333333333357 0.0000000000000000 F F F
0.7083333333333357 0.9583333333333357 0.0000000000000000 F F F
0.7083333333333357 0.2083333333333357 0.0000000000000000 F F F
0.9583333333333357 0.4583333333333357 0.0000000000000000 F F F
0.9583333333333357 0.7083333333333357 0.0000000000000000 F F F
0.9583333333333357 0.9583333333333357 0.0000000000000000 F F F
0.9583333333333357 0.2083333333333357 0.0000000000000000 F F F
0.2083333333333357 0.4583333333333357 0.0000000000000000 F F F
0.2083333333333357 0.7083333333333357 0.0000000000000000 F F F
0.2083333333333357 0.9583333333333357 0.0000000000000000 F F F
0.2083333333333357 0.2083333333333357 0.0000000000000000 F F F
0.3750000000000000 0.3750000000000000 0.9524831228666315 F F F
0.3750000000000000 0.6250000000000000 0.9524831228666315 F F F
0.3750000000000000 0.8750000000000000 0.9524831228666315 F F F
0.3750000000000000 0.1250000000000000 0.9524831228666315 F F F
0.6250000000000000 0.3750000000000000 0.9524831228666315 F F F
0.6250000000000000 0.6250000000000000 0.9524831228666315 F F F
0.6250000000000000 0.8750000000000000 0.9524831228666315 F F F
0.6250000000000000 0.1250000000000000 0.9524831228666315 F F F
0.8750000000000000 0.3750000000000000 0.9524831228666315 F F F
0.8750000000000000 0.6250000000000000 0.9524831228666315 F F F
0.8750000000000000 0.8750000000000000 0.9524831228666315 F F F
0.8750000000000000 0.1250000000000000 0.9524831228666315 F F F
0.1250000000000000 0.3750000000000000 0.9524831228666315 F F F
0.1250000000000000 0.6250000000000000 0.9524831228666315 F F F
0.1250000000000000 0.8750000000000000 0.9524831228666315 F F F
0.1250000000000000 0.1250000000000000 0.9524831228666315 F F F
0.2916666666666643 0.2916666666666643 0.9049662457332701 F F F
0.2916666666666643 0.5416666666666643 0.9049662457332701 F F F
0.2916666666666643 0.7916666666666643 0.9049662457332701 F F F
0.2916666666666643 0.0416666666666643 0.9049662457332701 F F F
0.5416666666666643 0.2916666666666643 0.9049662457332701 F F F
0.5416666666666643 0.5416666666666643 0.9049662457332701 F F F
0.5416666666666643 0.7916666666666643 0.9049662457332701 F F F
0.5416666666666643 0.0416666666666643 0.9049662457332701 F F F
0.7916666666666643 0.2916666666666643 0.9049662457332701 F F F
0.7916666666666643 0.5416666666666643 0.9049662457332701 F F F
0.7916666666666643 0.7916666666666643 0.9049662457332701 F F F
0.7916666666666643 0.0416666666666643 0.9049662457332701 F F F
0.0416666666666643 0.2916666666666643 0.9049662457332701 F F F
0.0416666666666643 0.5416666666666643 0.9049662457332701 F F F
0.0416666666666643 0.7916666666666643 0.9049662457332701 F F F
0.0416666666666643 0.0416666666666643 0.9049662457332701 F F F
0.2083333333333357 0.2083333333333357 0.8574493685999016 F F F
0.2083333333333357 0.4583333333333357 0.8574493685999016 F F F
0.2083333333333357 0.7083333333333357 0.8574493685999016 F F F
0.2083333333333357 0.9583333333333357 0.8574493685999016 F F F
0.4583333333333357 0.2083333333333357 0.8574493685999016 F F F
0.4583333333333357 0.4583333333333357 0.8574493685999016 F F F
0.4583333333333357 0.7083333333333357 0.8574493685999016 F F F
0.4583333333333286 0.9583333333333357 0.8574493685999016 F F F
0.7083333333333357 0.2083333333333357 0.8574493685999016 F F F
0.7083333333333357 0.4583333333333357 0.8574493685999016 F F F
0.7083333333333357 0.7083333333333357 0.8574493685999016 F F F
0.7083333333333286 0.9583333333333357 0.8574493685999016 F F F
0.9583333333333357 0.2083333333333357 0.8574493685999016 F F F
0.9583333333333357 0.4583333333333357 0.8574493685999016 F F F
0.9583333333333357 0.7083333333333357 0.8574493685999016 F F F
0.9583333333333357 0.9583333333333357 0.8574493685999016 F F F
0.1248753274088950 0.1248753274088950 0.8104723183369446 T T T
0.1250432471286117 0.3744669037981175 0.8104439099895814 T T T
0.1250432471286117 0.6254898490732638 0.8104439099895814 T T T
0.1248753274088950 0.8752493451822171 0.8104723183369446 T T T
0.3744669037981175 0.1250432471286117 0.8104439099895814 T T T
0.3750000000000000 0.3750000000000000 0.8101623576913966 T T T
0.3744669037981175 0.6254898490732638 0.8104439099895814 T T T
0.3741437503952032 0.8754281248023950 0.8106074679484812 T T T
0.6254898490732638 0.1250432471286117 0.8104439099895814 T T T
0.6254898490732638 0.3744669037981175 0.8104439099895814 T T T
0.6249056987982424 0.6249056987982424 0.8109175909471307 T T T
0.6249056987982424 0.8751886024035143 0.8109175909471307 T T T
0.8752493451822171 0.1248753274088950 0.8104723183369446 T T T
0.8754281248023950 0.3741437503952032 0.8106074679484812 T T T
0.8751886024035143 0.6249056987982424 0.8109175909471307 T T T
0.8754281248023950 0.8754281248023950 0.8106074679484812 T T T
0.0416666666666643 0.0416666666666643 0.7630725963525371 T T T
0.0423582548037352 0.2929829466331252 0.7632255425190596 T T T
0.0423342079934537 0.5413328960032665 0.7639551649056903 T T T
0.0423582548037352 0.7896587985631328 0.7632255425190596 T T T
0.2929829466331252 0.0423582548037352 0.7632255425190596 T T T
0.2921448802009530 0.2921448802009530 0.7631719676853072 T T T
0.2921448802009530 0.5407102395980798 0.7631719676853072 T T T
0.2929829466331252 0.7896587985631328 0.7632255425190596 T T T
0.5413328960032665 0.0423342079934537 0.7639551649056903 T T T
0.5407102395980798 0.2921448802009530 0.7631719676853072 T T T
0.5413328960032665 0.5413328960032665 0.7639551649056903 T T T
0.5423722548894060 0.7913138725552938 0.7643637592766812 T T T
0.7896587985631328 0.0423582548037352 0.7632255425190596 T T T
0.7896587985631328 0.2929829466331252 0.7632255425190596 T T T
0.7913138725552938 0.5423722548894060 0.7643637592766812 T T T
0.7913138725552938 0.7913138725552938 0.7643637592766812 T T T
0.9592783288347152 0.9592783288347152 0.7165014402517978 T T T
0.9592783288347081 0.2064433423305768 0.7165014402517978 T T T
0.9585429811439916 0.4580852035394961 0.7171342391736614 T T T
0.9585429811439987 0.7083718153165048 0.7171342391736614 T T T
0.2064433423305768 0.9592783288347152 0.7165014402517978 T T T
0.2080072672259897 0.2080072672259968 0.7154393564109170 T T T
0.2098510312054594 0.4575744843972738 0.7163450633092348 T T T
0.2080072672259968 0.7089854655480138 0.7154393564109170 T T T
0.4580852035394889 0.9585429811439987 0.7171342391736614 T T T
0.4575744843972738 0.2098510312054594 0.7163450633092348 T T T
0.4575744843972738 0.4575744843972738 0.7163450633092348 T T T
0.4580852035394961 0.7083718153165048 0.7171342391736614 T T T
0.7083718153165048 0.9585429811439987 0.7171342391736614 T T T
0.7089854655480138 0.2080072672259968 0.7154393564109170 T T T
0.7083718153165048 0.4580852035394961 0.7171342391736614 T T T
0.7083333333333357 0.7083333333333357 0.7173121617349102 T T T
0.9312769255399772 0.9312769255399772 0.3485062381768784 T T T
0.9312769255399772 0.2624461489200455 0.3485062381768784 T T T
0.9325931874642169 0.5962034062678949 0.3492759583004709 T T T
0.2624461489200455 0.9312769255399772 0.3485062381768784 T T T
0.2649012212242361 0.2649012212242361 0.3482925057539119 T T T
0.2649012212242361 0.5951975575515207 0.3482925057539119 T T T
0.5962034062678949 0.9325931874642169 0.3492759583004709 T T T
0.5951975575515207 0.2649012212242361 0.3482925057539119 T T T
0.5962034062678949 0.5962034062678949 0.3492759583004709 T T T
0.8194444444444429 0.8194444444444429 0.4120073015393046 F F F
0.8194444444444429 0.1527777777777786 0.4120073015393046 F F F
0.8194444444444429 0.4861111111111143 0.4120073015393046 F F F
0.1527777777777786 0.8194444444444429 0.4120073015393046 F F F
0.1527777777777786 0.1527777777777786 0.4120073015393046 F F F
0.1527777777777786 0.4861111111111143 0.4120073015393046 F F F
0.4861111111111143 0.8194444444444429 0.4120073015393046 F F F
0.4861111111111143 0.1527777777777786 0.4120073015393046 F F F
0.4861111111111143 0.4861111111111143 0.4120073015393046 F F F
0.7083333333333357 0.7083333333333357 0.4760019913289000 F F F
0.7083333333333357 0.0416666666666643 0.4760019913289000 F F F
0.7083333333333357 0.3750000000000000 0.4760019913289000 F F F
0.0416666666666643 0.7083333333333357 0.4760019913289000 F F F
0.0416666666666643 0.0416666666666643 0.4760019913289000 F F F
0.0416666666666643 0.3750000000000000 0.4760019913289000 F F F
0.3750000000000000 0.7083333333333357 0.4760019913289000 F F F
0.3750000000000000 0.0416666666666643 0.4760019913289000 F F F
0.3750000000000000 0.3750000000000000 0.4760019913289000 F F F
0.5972222222222214 0.5972222222222214 0.5399966811184953 F F F
0.5972222222222214 0.9305555555555571 0.5399966811184953 F F F
0.5972222222222214 0.2638888888888857 0.5399966811184953 F F F
0.9305555555555571 0.5972222222222214 0.5399966811184953 F F F
0.9305555555555571 0.9305555555555571 0.5399966811184953 F F F
0.9305555555555571 0.2638888888888857 0.5399966811184953 F F F
0.2638888888888857 0.5972222222222214 0.5399966811184953 F F F
0.2638888888888857 0.9305555555555571 0.5399966811184953 F F F
0.2638888888888857 0.2638888888888857 0.5399966811184953 F F F
0.4861111111111143 0.4861111111111143 0.6039913709080977 F F F
0.4861111111111143 0.8194444444444429 0.6039913709080977 F F F
0.4861111111111143 0.1527777777777786 0.6039913709080977 F F F
0.8194444444444429 0.4861111111111143 0.6039913709080977 F F F
0.8194444444444429 0.8194444444444429 0.6039913709080977 F F F
0.8194444444444429 0.1527777777777786 0.6039913709080977 F F F
0.1527777777777786 0.4861111111111143 0.6039913709080977 F F F
0.1527777777777786 0.8194444444444429 0.6039913709080977 F F F
0.1527777777777786 0.1527777777777786 0.6039913709080977 F F F
0.3750000000000000 0.3750000000000000 0.6709451889760421 T T T
0.3757686837881531 0.7078847317116083 0.6692728159340477 T T T
0.3757686837881531 0.0413465845002455 0.6692728159340477 T T T
0.7078847317116083 0.3757686837881531 0.6692728159340477 T T T
0.7083333333333357 0.7083333333333357 0.6672815507555968 T T T
0.7078847317116083 0.0413465845002455 0.6692728159340477 T T T
0.0413465845002455 0.3757686837881531 0.6692728159340477 T T T
0.0413465845002455 0.7078847317116083 0.6692728159340477 T T T
0.0416666666666643 0.0416666666666643 0.6705979807351840 T T T
0.0416666666666643 0.0416666666666643 0.3295224610717508 T T T
0.0415965476045645 0.3744525699724842 0.3318913105762326 T T T
0.0415965476045645 0.7089508824229516 0.3318913105762326 T T T
0.3744525699724842 0.0415965476045645 0.3318913105762326 T T T
0.3750000000000000 0.3750000000000000 0.3298742237993510 T T T
0.3744525699724842 0.7089508824229516 0.3318913105762326 T T T
0.7089508824229516 0.0415965476045645 0.3318913105762326 T T T
0.7089508824229516 0.3744525699724842 0.3318913105762326 T T T
0.7083333333333357 0.7083333333333357 0.3343592162472642 T T T
0.9305555555555571 0.9305555555555571 0.3960086290919023 F F F
0.9305555555555571 0.2638888888888857 0.3960086290919023 F F F
0.9305555555555571 0.5972222222222214 0.3960086290919023 F F F
0.2638888888888857 0.9305555555555571 0.3960086290919023 F F F
0.2638888888888857 0.2638888888888857 0.3960086290919023 F F F
0.2638888888888857 0.5972222222222214 0.3960086290919023 F F F
0.5972222222222214 0.9305555555555571 0.3960086290919023 F F F
0.5972222222222214 0.2638888888888857 0.3960086290919023 F F F
0.5972222222222214 0.5972222222222214 0.3960086290919023 F F F
0.8194444444444429 0.8194444444444429 0.4600033188815047 F F F
0.8194444444444429 0.1527777777777786 0.4600033188815047 F F F
0.8194444444444429 0.4861111111111143 0.4600033188815047 F F F
0.1527777777777786 0.8194444444444429 0.4600033188815047 F F F
0.1527777777777786 0.1527777777777786 0.4600033188815047 F F F
0.1527777777777786 0.4861111111111143 0.4600033188815047 F F F
0.4861111111111143 0.8194444444444429 0.4600033188815047 F F F
0.4861111111111143 0.1527777777777786 0.4600033188815047 F F F
0.4861111111111143 0.4861111111111143 0.4600033188815047 F F F
0.7083333333333357 0.7083333333333357 0.5239980086711000 F F F
0.7083333333333357 0.0416666666666643 0.5239980086711000 F F F
0.7083333333333357 0.3750000000000000 0.5239980086711000 F F F
0.0416666666666643 0.7083333333333357 0.5239980086711000 F F F
0.0416666666666643 0.0416666666666643 0.5239980086711000 F F F
0.0416666666666643 0.3750000000000000 0.5239980086711000 F F F
0.3750000000000000 0.7083333333333357 0.5239980086711000 F F F
0.3750000000000000 0.0416666666666643 0.5239980086711000 F F F
0.3750000000000000 0.3750000000000000 0.5239980086711000 F F F
0.5972222222222214 0.5972222222222214 0.5879926984606954 F F F
0.5972222222222214 0.9305555555555571 0.5879926984606954 F F F
0.5972222222222214 0.2638888888888857 0.5879926984606954 F F F
0.9305555555555571 0.5972222222222214 0.5879926984606954 F F F
0.9305555555555571 0.9305555555555571 0.5879926984606954 F F F
0.9305555555555571 0.2638888888888857 0.5879926984606954 F F F
0.2638888888888857 0.5972222222222214 0.5879926984606954 F F F
0.2638888888888857 0.9305555555555571 0.5879926984606954 F F F
0.2638888888888857 0.2638888888888857 0.5879926984606954 F F F
0.4857962518454938 0.4857962518454938 0.6520134262762931 T T T
0.4849632646245893 0.8200183676877012 0.6512691959389023 T T T
0.4857962518454938 0.1534074963090196 0.6520134262762931 T T T
0.8200183676877012 0.4849632646245893 0.6512691959389023 T T T
0.8200183676877012 0.8200183676877012 0.6512691959389023 T T T
0.8206112061993666 0.1521943969003203 0.6522258679919717 T T T
0.1534074963090195 0.4857962518454938 0.6520134262762931 T T T
0.1521943969003203 0.8206112061993666 0.6522258679919717 T T T
0.1521943969003203 0.1521943969003203 0.6522258679919717 T T T
After computing the electrostatic potential by setting WRT_POTENTIAL = hartree ionic, PREC = Accurate, and KSPACING = 0.2, we can run the Python script referenced above in Step 4.
For the Al(111)/Si(111) interface studied, we get the following results: [math]\displaystyle{ \bar{V}_{\mathrm{m}} = -0.694 }[/math] eV, [math]\displaystyle{ \bar{V}_{\mathrm{sc}} = +1.192 }[/math] eV, and [math]\displaystyle{ \Delta\bar{V} = -1.886 }[/math] eV.
Finally, we arrive at the Schottky barriers:
- [math]\displaystyle{ \varphi_{\mathrm{p}} = \Delta\bar{V} + E_{\mathrm{F}} - E_{\mathrm{VBM}} = -1.886 + 8.085 - 5.449 = 0.75 }[/math] eV,
which fits well to the experimental values that vary between 0.68 and 0.81 eV, depending on sample and experimental conditions[6], and
- [math]\displaystyle{ \varphi_{\mathrm{n}} = E_{\mathrm{g}} - \varphi_{\mathrm{p}} = 1.160 - 0.75 = 0.41 }[/math] eV.
Related tags and articles
LVHAR, WRT_POTENTIAL, BANDGAP, Computing the work function
References
- ↑ A. Baldereschi, S. Baroni, and R. Resta Band offsets in lattice-matched heterojunctions: A model and first-principles calculations for GaAs/AlAs, Phys. Rev. Lett. 61, 734 (1988).
- ↑ M. Peressi, M. Binggeli, and A. Baldereschi Band engineering at interfaces: theory and numerical experiments, J. Phys. D: Appl. Phys. 31, 1273 (1998).
- ↑ In VASP, the [math]\displaystyle{ \mathbf{G}=0 }[/math] Fourier component of the Hartree potential is set to zero by convention, so all single-particle eigenvalues are already referenced to the average electrostatic potential of their respective unit cell. As a consequence, [math]\displaystyle{ E_{\mathrm{F}} }[/math] and [math]\displaystyle{ E_{\mathrm{VBM}} }[/math] can be read directly from the bulk OUTCAR files without any further potential referencing, and only the interface calculation requires the electrostatic potential to be written out — to vaspout.h5 via WRT_POTENTIAL (recommended) or to LOCPOT via LVHAR (legacy).
- ↑ https://pymatgen.org/ (2022).
- ↑ https://wiki.fysik.dtu.dk/ase/ (2025).
- ↑ Ş. Altındal, H. Kanbur, A. Tataroğlu, and M.M. Bülbül, The barrier height distribution in identically prepared Al/p-Si Schottky diodes with the native interfacial insulator layer (SiO2), Physica B: Condensed Matter 399, 146-154 (2007).
